Summary and Schedule
The Machine Learning for Biologists (ML4Bio) workshop is aimed at biologists with no previous machine learning experience and minimal computational experience. This workshop is designed to teach machine learning concepts, not how to implement machine learning models. After the workshop, participants will be able to define machine learning concepts like samples, features, training set, validation set, test set, evaluation metrics, and model selection. Participants will be able to interactively develop an understanding of machine learning classifiers commonly used in biology like decision trees, random forests, logistic regression, and neural networks. The focus will be on problems in biology where machine learning is effectively used.
The ML4Bio workshop materials are still in development. Feedback is welcome in the GitHub issues.
Getting Started
This is not a programming workshop. You will not be taught how to code or implement machine learning models. This workshop can help you understand machine learning concepts. The goal is to provide you with resources to enable you to engage in conversations about machine learning, read articles that use machine learning, and understand the research methodology at a high level. The workshop uses point-and-click software without requiring any coding. This workshop focuses on supervised machine learning and classification.
To get started, follow the directions in the Setup page to access the required software and data for this workshop.
| Setup Instructions | Download files required for the lesson | |
| Duration: 00h 00m | 1. Introduction | What is machine learning? |
| Duration: 00h 10m | 2. Classifying T-cells | What are the steps in a machine learning workflow? |
| Duration: 00h 20m | 3. Evaluating a Model | How do you evaluate the performance of a machine learning model? |
| Duration: 00h 30m | 4. Decision Trees, Random Forests, and Overfitting | How do decision trees and random forests make decisions? |
| Duration: 00h 40m | 5. Logistic Regression, Artificial Neural Networks, and Linear Separability | What is linear separability? |
| Duration: 00h 50m | 6. Understanding Machine Learning Literature | How are machine learning workflows presented in research papers? |
| Duration: 01h 00m | 7. Conclusion and next steps | Where can you learn more about machine learning? |
| Duration: 01h 10m | Finish |
The actual schedule may vary slightly depending on the topics and exercises chosen by the instructor.
Software overview
Setup and launch the ml4bio software
Questions
- What do I need to run the ml4bio software?
- How do I download the ML4Bio workshop materials?
Objectives
- Provide instructions for installing the ml4bio software.
- Explain the ml4bio software environment
Overview
There are three main steps to prepare for the ML4Bio workshop:
- Download the workshop materials
- Install the Anaconda Python distribution
- Install the ml4bio software
See the troubleshooting if you run into problems during the installation. If you already have Python installed and do not want to use Anaconda, download the ML4Bio materials and proceed to the advanced instructions.
Download the ML4Bio materials
To download the ML4Bio materials, visit https://github.com/carpentries-incubator/ml4bio-workshop/.
Click the Code button followed by
Download ZIP.
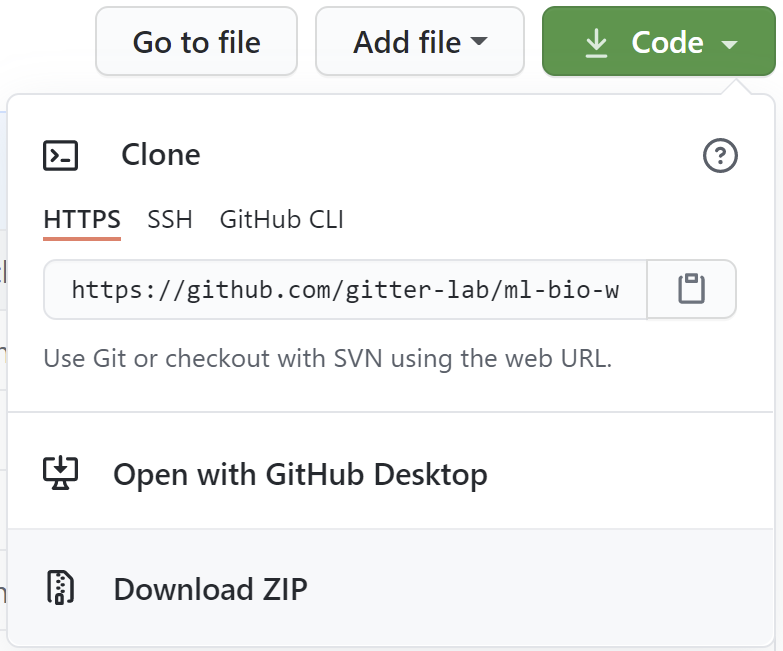
Save the file ml4bio-workshop-gh-pages.zip and then open
that location on your computer. Extract the zip file and open the folder
ml4bio-workshop-gh-pages, which has the same contents as https://github.com/carpentries-incubator/ml4bio-workshop/.
You are now ready to install the Python dependencies needed to run the
ml4bio software and follow the workshop exercises. You will also use the
datasets in the data subdirectory during the workshop. Take
note of the location of the ml4bio-workshop-gh-pages folder
so you can navigate to it during the workshop.
Install Python
ml4bio requires Python and several other Python packages. The easiest
way to install Python and the correct version of these packages is
through Anaconda, a Python
distribution. If you do not have Anaconda installed, please visit https://www.anaconda.com/download/ to download and
install the Python 3.x version (for example, 3.8). We recommend
letting the installer add Anaconda to your computer’s PATH
environment variable so that it is easily accessible from the command
line. This screenshot shows the PATH option in the
2019 version of the Anaconda Windows installer:
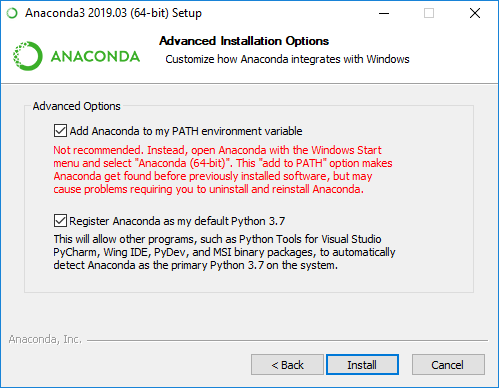
This will also make Anaconda your primary Python distribution. See the Carpentries Anaconda installation instructions for a step-by-step guide and video on how to install Anaconda for your operating system.
Launch the ml4bio software
After you install Anaconda, you will use installation scripts in the
scripts subdirectory of the
ml4bio-workshop-gh-pages directory to install the ml4bio
software. These are wrapper scripts that will run ml4bio inside a conda environment. If the
environment does not already exist, it will be created. This can take
5-10 minutes and requires internet connectivity to download the Python
packages.
- For Windows, launch the
install_launch_windows.batscript. You may need to run this script twice, once to install the software and again to launch it. - For Mac OS, launch the
install_launch_mac.commandscript. - For Linux, launch the
install_launch_linux.shscript.
To launch the correct script for your operating system, navigate to
the scripts subdirectory of the unzipped
ml4bio-workshop-gh-pages directory from the command line.
For Windows, launch the Anaconda Prompt (formerly Anaconda Command
Prompt) and then run the script:
- Start -> Type “Anaconda” -> Anaconda Prompt
- Navigate to the
ml4bio-workshop-gh-pages\scriptsdirectory from the command line using the commandcd <PATH_TO_ml4bio-workshop-gh-pages>\scripts(replace<PATH_TO_ml4bio-workshop-gh-pages>with the appropriate directory on your computer) - Type
install_launch_windows.bat-> Enter
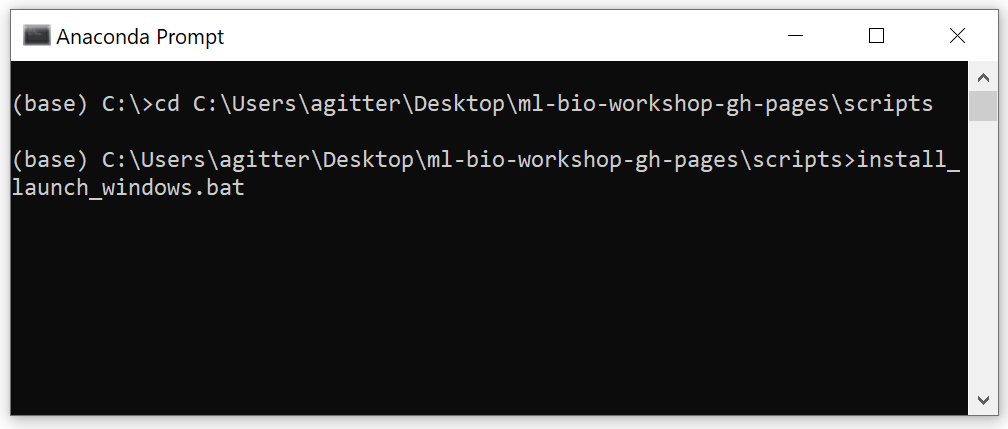
For Linux or Mac OS, open the terminal and navigate to the
ml4bio-workshop-gh-pages/scripts directory. Then, enter
./ followed by the name of the script for your operating
system without a space in between.
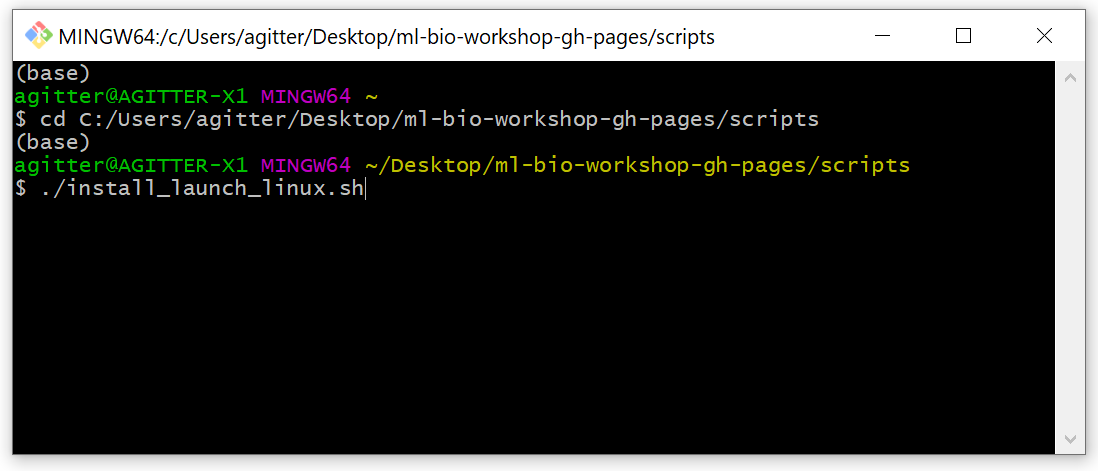
If the ml4bio software was successfully installed, you should see this graphical interface:
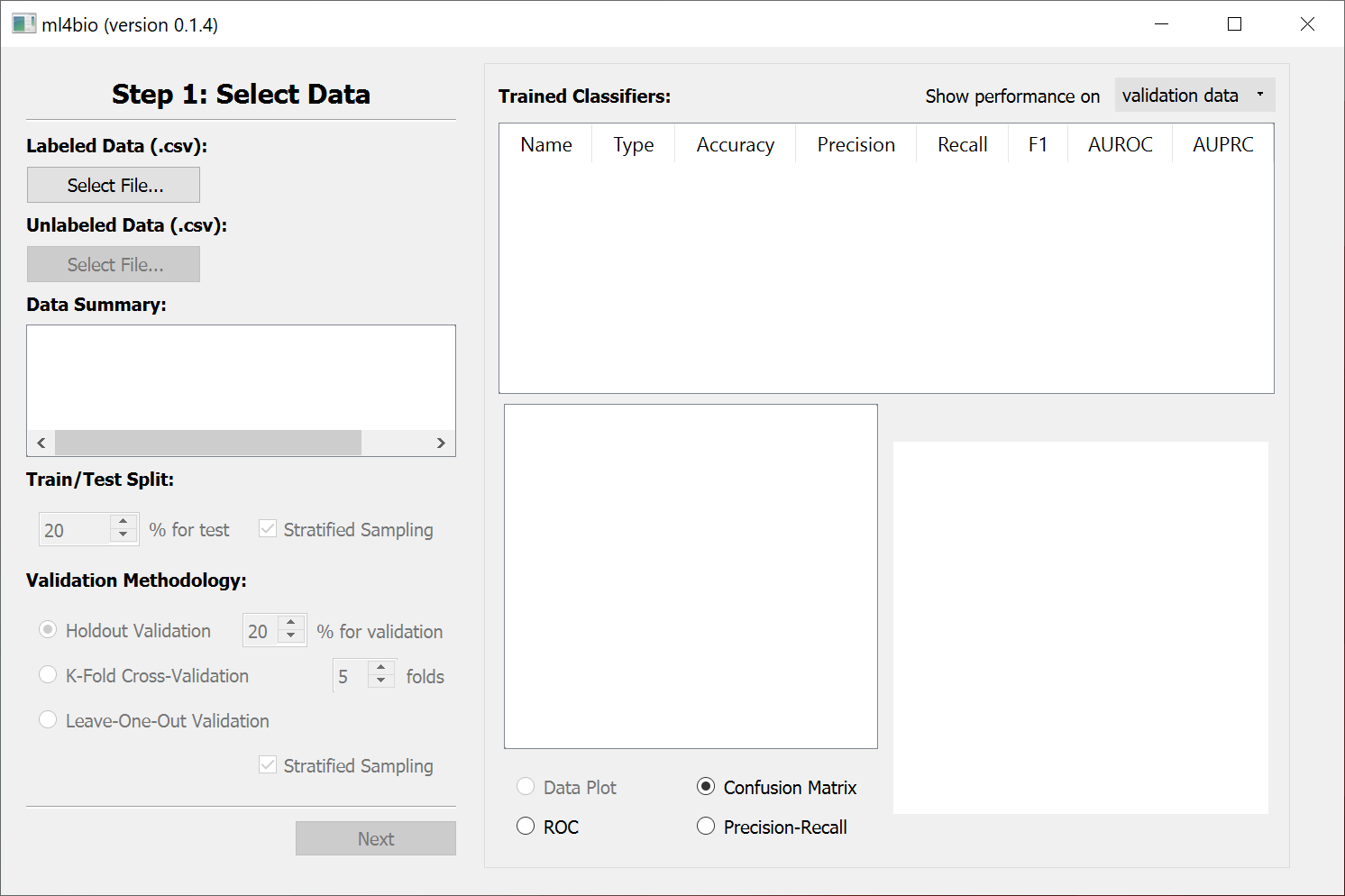
After you close the ml4bio software, you can run the same
install_launch script to relaunch it. The script will not
install anything new the second time you run it. It will use Anaconda to
open ml4bio.
A dedicated [lesson][episode-t-cells] will provide an introduction to the ml4bio software during the workshop. See the software environment details for more information about how the ml4bio software works.
Troubleshooting
You must extract the contents of the
ml4bio-workshop-gh-pages.zip workshop materials file. Even
though you may be able to browse the compressed directory to inspect the
files, the software installation will not work until the file is
unzipped.
If you did not add Anaconda to your PATH during
installation and would like to, follow these instructions for Windows
10:
- Start -> Type “Path” -> Edit environment variables for your account
- Path -> Edit -> New -> Browse -> Browse to the location where Anaconda was installed and select the Scripts subdirectory -> OK -> OK
When running the install_launch_windows.bat install
script, Windows may display a warning that the app is from an unknown
publisher and may be unsafe to run. This warning can be ignored.
See also known software warnings that can be safely ignored.
Updating ml4bio
New versions of the ml4bio software will be periodically released through PyPI. The release notes describe the changes in each new version. To install the latest version of ml4bio, run the appropriate update script for your operating system:
update_ml4bio_windows.batupdate_ml4bio_mac.commandupdate_ml4bio_linux.sh
Run these scripts in the same manner as the install scripts above.
Software environment details
Anaconda includes software that enables you to run Python programs as well as additional tools for managing software environments, programming in Python, and integrating code with textual descriptions and results in Jupyter notebooks. The software environments are managed by conda, one of the tools included with Anaconda. An environment is a collection of specific versions of Python packages. These are all stored in a directory that conda manages. Having multiple environments allows you to use different versions of the same package for different projects.
The ml4bio install scripts create a new conda environment. This
environment, which is named ml4bio, contains the latest
version of the ml4bio Python package as well as suitable
versions of other Python packages that it requires. The
ml4bio code may be incompatible with older or newer
versions of the Python packages it uses. The environment makes it easy
for you use a collection of Python packages that work together.
The most important required Python package that ml4bio uses is called scikit-learn. This is a popular general purpose machine learning package. When you use the ml4bio graphical interface, it calls functions in scikit-learn to train classifiers and make predictions.
Advanced users
Advanced users who already have Python installed can install the required packages through pip. Then launch ml4bio
from the command line with the command ml4bio.
