Content from Introduction to Python
Last updated on 2026-02-20 | Edit this page
Overview
Questions
- How can I store information?
- How can I perform actions?
Objectives
- Assign values to variables.
- Run built-in functions.
- Import libraries.
Using Python
In the following episodes, we will learn the fundamental knowledge to read and run the Python programming language. To run Python commands, we will use a Jupyter Notebook, an interactive platform to run code, write text, and see your results, such as plots. Follow the instructions in the Setup page to open a Notebook.
Variables
In Python, you store information in variables, which have a name and an assigned value.
PYTHON
variable_name = "value" # Here we assign the value to the variable name
variable_name # Here we ask to see the valueOUTPUT
'value'The values can be the result of operations.
OUTPUT
'aLongerValue'Until now, the data type we have been using is string
(str). We saw that summing strings results in a longer
combined string. If we work with numerical values the result will be the
mathematical one.
OUTPUT
300OUTPUT
600We can also perform operations with variables that are already set.
OUTPUT
1000And if we want to be more efficient, we can assign values to more than one variable in the same line.
OUTPUT
'blue'OUTPUT
'big'Built-in functions and libraries
Python has basic functions to carry out specific actions. In Python,
the arguments of the functions go inside parentheses, and different
functions take different kinds and numbers of arguments. The function
len() takes one argument and outputs its length. In the
case of a string, the length is the number of characters.
OUTPUT
12ERROR
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
/tmp/ipykernel_55271/3566361226.py in <module>
1 a_number = 300
----> 2 len(a_number)
TypeError: object of type 'int' has no len()The len() function can give us the number of characters
in a string, but it can not do anything with a number (that is of the
type int, meaning integer).
Here, you can check the built-in functions of the Python language.
For extra functionalities, there are libraries, which are collections of functions and data structures (see the next episode) specialized in a common set of tasks.
Pandas is an indispensable library specialized in working with tabular data. To make it accessible, you need to import it. There are several ways to import libraries.
Import a library while establishing an alias to access the functions using the alias.
Import a library without an alias. You need to access the functions with the complete name of the library.
Import only certain functions from a library. You don’t need to specify a name or alias when calling the function.
Exercise 1(Begginer): Creating variables and using functions
Create three variables with the common name of your favorite bird as the variable name and the scientific name as the value. Use built-in functions to get the length of the scientific name that has the maximum length.
- Variables are Python objects that store information when you assign it to them.
- Python has built-in functions that are always accessible.
- Libraries give you access to more functions.
Content from Data Structures
Last updated on 2026-02-20 | Edit this page
Overview
Questions
- How can I store information in Python?
- How can I access the elements of an array or a dictionary?
- How can I efficiently manage tabular data in Python?
Objectives
- Understand fundamental data structures such as lists, arrays, dictionaries, and Pandas dataframes.
- Efficient data manipulation, selection and filtering.
- Tabular data management with Pandas.
- Read data from a URL.
List
A list is an indexed, ordered, changeable sequence of elements that
can be of different types and it allows duplicate values. They are
defined using square brackets []:
OUTPUT
['bacteria', 'archea', 'fungus', 4]To access elements in a list, we can use their index.
OUTPUT
bacteriaWe can also access elements in the list using negative indices, which count from the end of the list towards the beginning.
OUTPUT
4You can add elements to a list using the append() method
to add an element to the end of the list.
OUTPUT
['bacteria', 'archea', 'fungus', 4, 'animal']You can also extend a list by adding all the elements from another
list using the extend() method.
OUTPUT
['bacteria', 'archea', 'fungus', 4, 'animal', 1, 2, 3]You can remove elements from a list using the word del
followed by the index of the element you want to remove.
OUTPUT
['bacteria', 'archea', 'fungus', 'animal', 1, 2, 3]You can also use the remove() method to remove a
specific element by its value.
OUTPUT
['bacteria', 'archea', 'fungus', 1, 2, 3]You can concatenate two lists using the + operator or
the extend() method.
PYTHON
mylist1 = ['bacteria', 'archea', 'fungus']
mylist2 = ['animal', 'algae', 'plant']
newlist = mylist1 + mylist2
print(newlist)OUTPUT
['bacteria', 'archea', 'fungus', 'animal', 'algae', 'plant']The sort() method sorts the elements of the list in
ascending order by default. All elements need to be of the same type. If
we try to sort the elements of the list thislist, we will
get an error.
OUTPUT
TypeError: '<' not supported between instances of 'int' and 'str'OUTPUT
['bacteria', 'archea', 'fungus', 'animal', 'algae', 'plant']To sort them in descending order:
OUTPUT
['plant', 'algae', 'animal', 'fungus', 'archea', 'bacteria']Consider that the sort() method modifies the original
list and does not return any value. To sort a list without modifying the
original, you can use the sorted() function, which returns
a new sorted list.
Select elements of a list with slices
Another way to access list data in Python is by using the “slice”
method: list[start:end]. This slice includes the elements whose indices
are in the range from start to end - 1:
- start: It is the index from which we start including elements in the slice.
- end: It is the index up to which we include elements in the slice, but not including the element at this index.
For example, if we want to get a slice of newlist from
index 1 to index 4, we would use the notation newlist[1:4]
and we will get a slice that includes elements from index 1 (inclusive)
to index 4 (exclusive).
OUTPUT
['algae', 'animal', 'fungus']Another syntax of “slices” in Python with the notation list[start🔚step] refers to the technique for selecting a subset of elements from a list using three parameters:
- start: Indicates the index from which we start including elements in the slice.
- end: Indicates the index up to which we include elements in the slice, excluding the element at this index.
- step: Indicates the size of the step or increment between the selected elements. That is, how many elements we skip in each step.
If we run newlist[::2], we get a slice that includes
elements of the newlist, but selecting every second
element. It’s like starting from the beginning of the list, ending at
the end, and selecting every second element.
OUTPUT
['plant', 'animal', 'archea']You can remove “TCGA” from the list using the remove() method:
To add “CATG” to the list, you can use the append() method:
To find the sequence at index 2, you can access the element at that index using indexing.
sequence_at_index_2 = genes[2]
print(sequence_at_index_2)OUTPUT
CGTADictionary
Dictionaries (dict) are collections of key-value pairs. Each element
in the dictionary has a key and an associated value. They are defined
using curly braces {} and separating the keys and values with colons
:. Dictionaries in Python are mutable, meaning you can add,
modify, and delete elements as needed.
OUTPUT
{'brand': 'Ford', 'model': 'Mustang', 'year': 1964}You can access the values of the dictionary using their keys.
OUTPUT
FordOUTPUT
1964If you try to access a key that does not exist in the dictionary, Python will raise a KeyError.
OUTPUT
KeyError: 'color'You can add a new key-value pair to the dictionary by simply assigning a value to a new key.
OUTPUT
{'brand': 'Ford', 'model': 'Mustang', 'year': 1964, 'color': 'red'}You can modify the value associated with an existing key in the dictionary.
OUTPUT
{'brand': 'Ford', 'model': 'Mustang', 'year': 2022, 'color': 'red'}You can delete an item from the dictionary using the del
word.
OUTPUT
{'brand': 'Ford', 'model': 'Mustang', 'year': 2022}You can also use the pop() method to remove an item and
return its value.
OUTPUT
redOUTPUT
{'brand': 'Ford', 'model': 'Mustang', 'year': 2022}Reserved methods for dictionaries include keys(),
values(), and items(). These methods provide
convenient ways to access different aspects of a dictionary:
-
keys(): This method returns a view object that displays a list of all the keys in the dictionary. It allows you to iterate over the keys or convert them into a list if needed.
OUTPUT
dict_keys(['brand', 'model', 'year'])-
values(): This method returns a view object that displays a list of all the values in the dictionary. Similar tokeys(), it allows iteration over the values or conversion into a list.
OUTPUT
dict_values(['Ford', 'Mustang', 2022])-
items(): This method returns a view object that displays a list of tuples, where each tuple contains a key-value pair from the dictionary. It’s useful for iterating over both keys and values simultaneously, or for converting the dictionary into a list of key-value pairs.
OUTPUT
dict_items([('brand', 'Ford'), ('model', 'Mustang'), ('year', 2022)])Modify a dictionary
You can modify (update) a dictionary using the update()
method. When you call the update() method on a dictionary,
you pass another dictionary as an argument. The method then iterates
over the key-value pairs in the second dictionary and adds them to the
first dictionary. If any keys in the second dictionary already exist in
the first dictionary, their corresponding values are updated to the new
values.
OUTPUT
{'brand': 'Ford', 'model': 'Focus', 'year': 2022, 'price': 30000}To add new entries to our dictionary, we use the
append() method.
PYTHON
thisdict = {
'brand': ['Ford', 'Toyota'],
'model': ['Mustang', 'Corolla'],
'year': [1964, 2020],
'color': ['red', 'blue'],
'price': [15000, 20000]
}
# Add a new brand, model, year, color, and price
new_brand = 'Honda'
new_model = 'Civic'
new_year = 2019
new_color = 'green'
new_price = 18000
thisdict['brand'].append(new_brand)
thisdict['model'].append(new_model)
thisdict['year'].append(new_year)
thisdict['color'].append(new_color)
thisdict['price'].append(new_price)
# Print the updated dictionary
print(thisdict)OUTPUT
{'brand': ['Ford', 'Toyota', 'Honda'], 'model': ['Mustang', 'Corolla', 'Civic'], 'year': [1964, 2020, 2019], 'color': ['red', 'blue', 'green'], 'price': [15000, 20000, 18000]}One way to get the model and year values for each brand is as follows.
PYTHON
# Get the year and model of each car directly from the dictionary
ford_year = thisdict['year'][thisdict['brand'].index('Ford')]
ford_model = thisdict['model'][thisdict['brand'].index('Ford')]
toyota_year = thisdict['year'][thisdict['brand'].index('Toyota')]
toyota_model = thisdict['model'][thisdict['brand'].index('Toyota')]
# Print the year and model of each car
print("Ford's Year:", ford_year)
print("Ford's Model:", ford_model)
print("Toyota's Year:", toyota_year)
print("Toyota's Model:", toyota_model)OUTPUT
Ford's Year: 1964
Ford's Model: Mustang
Toyota's Year: 2020
Toyota's Model: CorollaExercise 2(Begginer): Manipulating Dictionaries
Suppose that you have the following dictionary:
genes_dict = {"gene1": { "name": "BRCA1", "start_position": 43044295, "end_position": 43125483},
"gene2": { "name": "TP53", "start_position": 7571720, "end_position": 7588830},
"gene3": { "name": "EGFR", "start_position": 55086715, "end_position": 55225454}}
~~~
{: .language-python}
Which command correctly adds a new gene "gene4" with its associated information to an existing genes dictionary in Python?
a) `genes_dict.add("gene4", {"name": "GENE4", "start_position": 123456, "end_position": 234567})`
b) `genes_dict["gene4"] = {"name": "GENE4", "start_position": 123456, "end_position": 234567}`
c) `genes_dict.append("gene4": {"name": "GENE4", "start_position": 123456, "end_position": 234567})`
d) `genes_dict.insert(4, {"name": "GENE4", "start_position": 123456, "end_position": 234567})`
> ## Solution
>
> The correct answer is option b).
> It uses the indexing syntax to add a new key-value pair to the genes_dict dictionary, where the key is "gene4" and the value is a dictionary representing the gene's characteristics.
>
{: .solution}Array
An method for creating and manipulating multidimensional arrays is by using NumPy. NumPy serves as a foundational library for scientific computing in Python, offering robust support for multidimensional arrays and an extensive array of mathematical functions for efficient memory utilization and rapid numerical operations.
To import it, we use the following.
To create a one-dimensional array, we use the following.
OUTPUT
[1 2 3 4 5 6]In NumPy, there are functions to create arrays of zeros or ones. To
create an array filled with zeros, you can use
np.zeros(shape), where shape is the desired shape of the
array:
OUTPUT
[0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]To create an array filled with ones, you can use
np.ones(shape).
OUTPUT
[1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]Other ways to create arrays and multidimensional arrays
Another way to create arrays is by using sequences with
arange(), for example:
OUTPUT
array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])Or using a notation similar to slices:
arange(start, stop, step):
OUTPUT
array([1, 2, 3, 4, 5, 6, 7, 8, 9])We can also create them using random values:
OUTPUT
array([9, 6, 0, 2, 7])Multidimensional arrays can be created in various ways by specifying the dimensions of each dimension. For example, to create a 2-dimensional array filled with zeros, ones, or random values:
PYTHON
array_zeros = np.zeros((3,3))
array_ones = np.ones((3,3))
array_rand = np.random.randint(0,10,(3,3))
print(array_zeros)
print(array_ones)
print(array_rand)OUTPUT
[[0. 0. 0.]
[0. 0. 0.]
[0. 0. 0.]]
[[1. 1. 1.]
[1. 1. 1.]
[1. 1. 1.]]
[[4 5 4]
[1 7 8]
[7 9 9]]Another way to create a two dimensional array is:
OUTPUT
[[1 2 3]
[4 5 6]
[7 8 9]]The expression len(array) returns the length of the
array, which corresponds to the number of elements in the array. For a
one-dimensional array, this is the number of elements it contains. For a
two-dimensional array, it is the number of rows in the array.
OUTPUT
10The NumPy library also allows us to load databases using
loadtxt. We will use a toy dataset to learn how to import a
csv file into a numpy array. The request
module is used in Python to make HTTP requests to web servers.
PYTHON
import requests
# The url to the csv file
url = "https://raw.githubusercontent.com/carpentries-incubator/pangenomics/gh-pages/files/spiral_2d.csv"
# Get the content of the file
response = requests.get(url)
content = response.text
# Load the data into a NumPy array
data = np.loadtxt(content.splitlines(), delimiter=' ')
dataOUTPUT
array([[ 2.728, 6.513],
[ 3.776, 26.114],
[ 5.595, 38.47 ],
...,
[2003.23 , 1849.23 ],
[2003.95 , 1928.41 ],
[2004.09 , 1966.77 ]])Exercise 3(Begginer): Manipulating arrays
Suppose you have a the following array dna_array containing DNA sequences as strings.
PYTHON
dna_sequences = ["AGCT", "TCGA", "ATCG", "CGTA", "GATTACA"]
dna_array = np.array(dna_sequences)
print(dna_array)You want to extract the sequences that meet a specific condition.
Which NumPy function would you use to extract DNA sequences from
dna_array that contain “AT”?
np.extract(dna_array == 'AT', dna_array)np.where(dna_array == 'AT')np.extract(np.char.startswith(dna_array, 'AT'), dna_array)dna_array[np.char.count(dna_array, 'AT') > 0]
The correct is d).
DataFrame
As we see in the first episode, Pandas is an indispensable library
specialized in working with tabular data or a DataFrame. We will import
the Pandas with its usual alias pd.
One form to create a DataFrame is with a dictionary. We will use the
dictionary thisdict and then we will convert it to a
dataframe.
OUTPUT
{'brand': ['Ford', 'Toyota', 'Honda'], 'model': ['Mustang', 'Corolla', 'Civic'], 'year': [1964, 2020, 2019], 'color': ['red', 'blue', 'green'], 'price': [15000, 20000, 18000]}We will add a row name with index.
OUTPUT
brand model year color price
C1 Ford Mustang 1964 red 15000
C2 Toyota Corolla 2020 blue 20000
C3 Honda Civic 2019 green 18000We can explore the dataframe using different methods, such as head(), tail(), info(), describe(), etc. For example:
OUTPUT
brand model year color price
C1 Ford Mustang 1964 red 15000
C2 Toyota Corolla 2020 blue 20000OUTPUT
brand model year color price
C2 Toyota Corolla 2020 blue 20000
C3 Honda Civic 2019 green 18000OUTPUT
year price
count 3.000000 3.000000
mean 2001.000000 17666.666667
std 32.046841 2516.611478
min 1964.000000 15000.000000
25% 1991.500000 16500.000000
50% 2019.000000 18000.000000
75% 2019.500000 19000.000000
max 2020.000000 20000.000000OUTPUT
<class 'pandas.core.frame.DataFrame'>
Index: 3 entries, C1 to C3
Data columns (total 5 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 brand 3 non-null object
1 model 3 non-null object
2 year 3 non-null int64
3 color 3 non-null object
4 price 3 non-null int64
dtypes: int64(2), object(3)
memory usage: 144.0+ bytes
NonePandas provides different methods for indexing and selecting data
from DataFrames. You can use iloc[] for integer-based
indexing and loc[] for label-based indexing. For
example:
- Select columns by name:
OUTPUT
C1 Mustang
C2 Corolla
C3 Civic
Name: model, dtype: object- Select rows by index using
iloc[]
OUTPUT
brand Toyota
model Corolla
year 2020
color blue
price 20000
Name: C2, dtype: object- Select rows by index name using
loc[]
OUTPUT
brand Ford
model Mustang
year 1964
color red
price 15000
Name: C1, dtype: object- Select rows and columns with
iloc[]
OUTPUT
brand model
C1 Ford Mustang
C2 Toyota Corolla- Select multiple columns.
OUTPUT
model year
C1 Mustang 1964
C2 Corolla 2020
C3 Civic 2019We can filter the data by some conditions. For example:
OUTPUT
brand model year color price
C2 Toyota Corolla 2020 blue 20000
C3 Honda Civic 2019 green 18000We can modify DataFrames by adding or removing rows and columns, updating values, and performing various data transformations. For example:
OUTPUT
brand model year color price mileage
C1 Ford Mustang 1964 red 15000 150000
C2 Toyota Corolla 2020 blue 20000 5000
C3 Honda Civic 2019 green 18000 30000OUTPUT
brand model year price mileage
C1 Ford Mustang 1964 15000 150000
C2 Toyota Corolla 2020 20000 5000
C3 Honda Civic 2019 18000 30000OUTPUT
brand model year price mileage
C1 Ford Mustang 1964 15000 150000
C2 Toyota Corolla 2020 20000 25000
C3 Honda Civic 2019 18000 30000Just like with numpy, we can read CSV files that are stored on our computer or on the internet.
PYTHON
# Read the url database
url = "https://raw.githubusercontent.com/carpentries-incubator/pangenomics/gh-pages/files/familias_minis.csv"
df_genes = pd.read_csv(url, index_col=0)
df_genes.head(5)OUTPUT
g_A909 g_2603V g_515 g_NEM316
A909|MGIDGNCP_01408 A909|MGIDGNCP_01408 2603V|GBPINHCM_01420 515|LHMFJANI_01310 NEM316|AOGPFIKH_01528
A909|MGIDGNCP_00096 A909|MGIDGNCP_00096 2603V|GBPINHCM_00097 515|LHMFJANI_00097 NEM316|AOGPFIKH_00098
A909|MGIDGNCP_01343 A909|MGIDGNCP_01343 NaN NaN NEM316|AOGPFIKH_01415
A909|MGIDGNCP_01221 A909|MGIDGNCP_01221 NaN 515|LHMFJANI_01130 NaN
A909|MGIDGNCP_01268 A909|MGIDGNCP_01268 2603V|GBPINHCM_01231 515|LHMFJANI_01178 NEM316|AOGPFIKH_01341 Exercise 4(Advanced): Manipulating dataframes
Use the dataframe df_genes and add a column that counts
how many genes are in each row, for example, the first row has 4 genes
but the third row has only two.
Another data structures: Sets and Tuples
Tuple
Tuples are indexed and ordered sequences, they are immutable, meaning
they cannot be modified after creation. The elements of a tuple can be
numbers, strings, or combinations of both types. A tuple is defined
using parentheses ():
OUTPUT
('bacteria', 'archea', 'fungus', 3)To access the elements of a tuple, we can use its index. In Python, indexing typically starts at 0. This means that the first element in a sequence (such as a list, tuple, or string) has an index of 0, the second element has an index of 1, and so on.
OUTPUT
bacteriaWe can also access tuple elements using negative indices, which count from the end of the tuple towards the beginning.
OUTPUT
3Set
Sets are unindexed, unordered collections of unique element, duplicates are not allowed. The sets can contain different data types. They are defined using curly braces {}:
OUTPUT
{'archea', 3, 'fungus', 'bacteria'}Sets in Python are mutable, meaning you can add and remove elements,
but you cannot directly modify existing elements. To add elements to a
set, you can use the add() method.
OUTPUT
{3, 'fungus', 'bacteria', 'animal', 'archea'}To remove an element from a set, you can use the
remove().
OUTPUT
{'fungus', 'bacteria', 'animal', 'archea'}Sets are useful when you need to store a collection of unique elements and perform efficient set operations such as removing duplicates and comparing collections.
- Gain familiarity with fundamental data structures such as tuples, sets, lists, dictionaries, and arrays.
- Develop skills in manipulating data by accessing, modifying, and filtering elements within data structures.
- Learn to work with tabular data using Pandas DataFrames, including loading and exploring data.
Content from Functions
Last updated on 2026-02-20 | Edit this page
Overview
Questions
- How can evaluate a condition?
- How can I repeat an action?
- How can create my custom functions?
Objectives
- Use the
ifconditional to decide an action - Use the
forloop to iterate actions - Create your functions
If statements
The if statement is a conditional statement in Python
that allows us to execute a block of code only if a certain condition is
true. It follows this syntax:
Let’s look at whether the two characters representing nucleotides are equal.
The else conditional executes a block when the condition in the if is not true.
PYTHON
char1='A'
char1='G'
if char1 == char2:
print( char1,"equal", char2)
else:
print( char1,"is not equal", char2)OUTPUT
A is not equal GYou can add as many instructions inside the if as you
want by indenting the sentences. Just remember always to keep the same
amount of characters in the indentation.
For loops
The for loop is used in Python to iterate over a sequence (such as a list, tuple, string, etc.) and execute a block of code for each element in the sequence. It follows this syntax:
Functions
Apart from the built-in functions and the library functions, you can make your functions, first, you define them, and then you use them. To define a function you need to give it a name, say what information you need to feed to it, and what output you want.
Here we will wrap the conditional to decide if two characters are equal in the function equal_chars. First, we defined the function and then we called it.
PYTHON
def equal_chars(character1, character2):
value=""
if character1 == character2:
value="equal"
else:
value="Not equal"
return valueAfter we create our first function, Let’s create a hamming distance function for two strings. The function hamming_distance() takes two parameters: str1 and str2, the two strings for which the Hamming distance is calculated.
First, we check if the lengths of the two strings are equal. If they are not equal, we raise a ValueError because the Hamming distance is only defined for strings of equal length. We initialize a variable distance to store the Hamming distance. We then iterate over the characters of the two strings using the zip() function, which pairs the corresponding elements of the two strings together. For each pair of characters, if they are different, we increment the distance variable. Finally, we return the calculated Hamming distance. You can use this function to calculate the Hamming distance between any two strings in Python.
PYTHON
def hamming_distance(str1, str2):
# Check if the strings have equal length
if len(str1) != len(str2):
raise ValueError("Strings must have equal length")
# Initialize the Hamming distance to 0
distance = 0
# Iterate over the characters of the strings
for char1, char2 in zip(str1, str2):
# If the characters are different, increment the distance
if char1 != char2:
distance += 1
# Return the calculated Hamming distance
return distanceNow lets add a multiline comment documenting the parameters needed for the fucntion.
PYTHON
def hamming_distance(str1, str2):
"""
Calculate the Hamming distance between two strings.
Parameters:
str1 (str): First string
str2 (str): Second string
Returns:
int: Hamming distance between the two strings
"""
# Check if the strings have equal length
if len(str1) != len(str2):
raise ValueError("Strings must have equal length")
# Initialize the Hamming distance to 0
distance = 0
# Iterate over the characters of the strings
for char1, char2 in zip(str1, str2):
# If the characters are different, increment the distance
if char1 != char2:
distance += 1
# Return the calculated Hamming distance
return distanceExercise 2(Intermediate): Sort the function to count the nucleotides in a string
Combining the data types you just learned, it is possible to automatize the function even more. Suppose that “population” is a numpy ndarray with genomes as rows. Genomes are represented with only two characters: 1 and 0.
Exercise 3(Intermediate): Sort the comments to document a Hamming distance function for matrixes
PYTHON
# The Hamming distance is multiplied by the number of genes to convert it into an absolute distance
# Number of genomes
# Calculate the Hamming distance between each pair of genomes
# Saving the distance in the matrix
# Create an empty matrix for Hamming distances
def calculate_hamming_matrix(population):
# COMMENT 1
num_genomes = population.shape[0]
# COMMENT 2
hamming_matrix = np.zeros((num_genomes, num_genomes), dtype=int)
# COMMENT 3
for i in range(num_genomes):
for j in range(i+1, num_genomes): # j=i+1 to avoid calculating the same distance twice
#COMMENT 4
distance = hamming(population[i], population[j]) * len(population[i])
hamming_matrix[i, j] = distance # COMMENT 5
hamming_matrix[j, i] = distance # The matrix is symmetric
return hamming_matrixPYTHON
def calculate_hamming_matrix(population):
# Number of genomes
num_genomes = population.shape[0]
# Create an empty matrix for Hamming distances
hamming_matrix = np.zeros((num_genomes, num_genomes), dtype=int)
# Calculate the Hamming distance between each pair of genomes
for i in range(num_genomes):
for j in range(i+1, num_genomes): # j=i+1 to avoid calculating the same distance twice
# The Hamming distance is multiplied by the number of genes to convert it into an absolute distance
distance = hamming(population[i], population[j]) * len(population[i])
hamming_matrix[i, j] = distance
hamming_matrix[j, i] = distance # The matrix is symmetric
return hamming_matrix- The conditional
ifevaluates a statement and performs an action - The for loop repeats an action a predetermined number of times
- Custom functions can be defined with
def
Content from Plotting
Last updated on 2026-02-20 | Edit this page
Overview
Questions
- How can I plot histograms
- How can I plot a graph with vertex and edges
Objectives
- Create a graph with matplotlib
- Create a graph with NetworkX
Python has several libraries that plot and visualize data. NetworkX is a library for working with graph data structures and algorithms, Plotly is a library for creating interactive and publication-quality plots, and Matplotlib is a comprehensive plotting library for creating static and interactive visualizations in Python. Each of these libraries serves different purposes and can be used for various data visualization tasks depending on the requirements and preferences of the user.
Plot with matplotlib
First, we import the matplotlib.pyplot module, which provides a MATLAB-like plotting interface.
Now, we defined some sample data for which we wanted to create a histogram.
OUTPUT
[1, 2, 2, 3, 3, 3, 4, 4, 4, 4, 5, 5, 5, 5, 5]
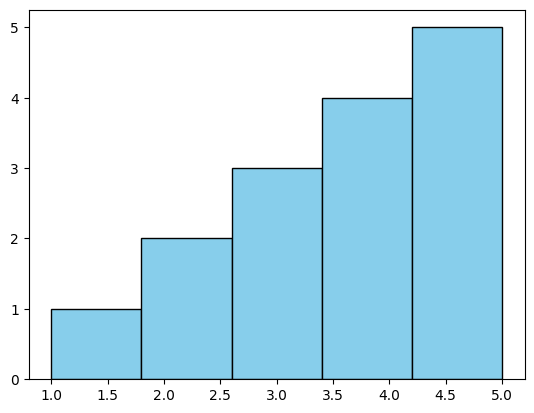
We use the plt.hist() function to create the histogram. We pass the data as the first argument, specify the number of bins using the bins parameter, and optionally specify the color of the bars and their edges using the color and edgecolor parameters, respectively.
OUTPUT
(array([1., 2., 3., 4., 5.]),
array([1. , 1.8, 2.6, 3.4, 4.2, 5. ]),
<BarContainer object of 5 artists>)We add labels to the x-axis and y-axis using plt.xlabel() and plt.ylabel(), and we add a title to the plot using plt.title().
PYTHON
plt.xlabel('Value')
plt.ylabel('Frequency')
plt.title('Histogram of Sample Data')
plt.hist(data, bins=5, color='skyblue', edgecolor='black')OUTPUT
(array([1., 2., 3., 4., 5.]),
array([1. , 1.8, 2.6, 3.4, 4.2, 5. ]),
<BarContainer object of 5 artists>)Notice that if you are not in colab or Jupyter Notebook, but in stand-alone Python you will need plt.show() to display the plot.
In this example, we produced a simple histogram of the sample data with five bins, with each bin representing the frequency of values falling within its range. The bars were colored sky blue with black edges for better visibility.
Exercise 1(Begginer): The nucleotide frequency of a DNA sequence
In the DNA sequence stored in the string dna_sequence, we want to
graph the frequency of each nucleotide. Sort and fill in the blanks in
the following code to get the frequency plot. Notice that we are using
the function nucleotide_counts, which we constructed in the previous
episode
 Figure 5. Histogram of the nucleotide frequency on a DNA
sequence.
Figure 5. Histogram of the nucleotide frequency on a DNA
sequence.
PYTHON
nucleotide_counts = calculate_nucleotide_frequency(dna_sequence)
nucleotides = list(nucleotide_counts.keys())
frequencies = list(nucleotide_counts.values())
plt.xlabel('Nucleotide')
plt.ylabel('Frequency')
plt.title('Nucleotide Frequency Histogram')
plt.bar(nucleotides, frequencies, color='skyblue')Creating graphs with NetworkX and Plotly
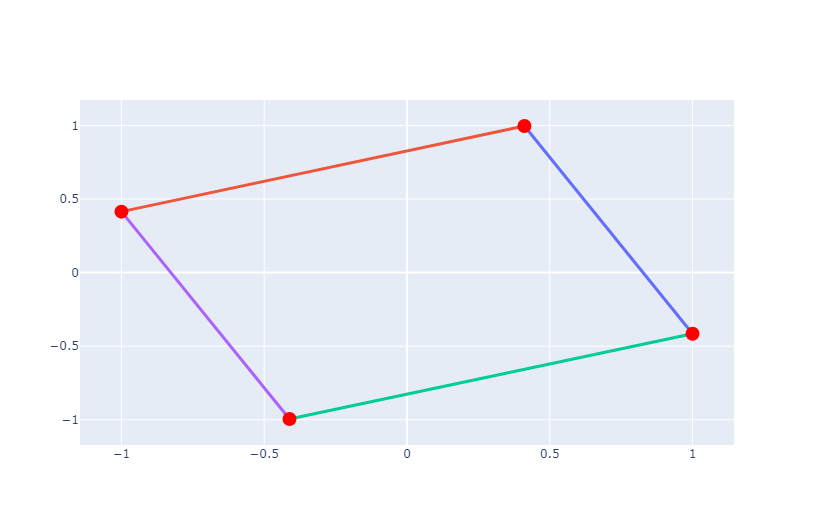
Let’s create a simple graph with NetworkX and visualize it using Plotly. In this example, we’ll create a graph with four nodes and four edges. Nodes are represented as red markers, and edges are represented as black lines.
We create a simple undirected graph G.
We define positions for nodes using a spring layout algorithm (spring_layout). This assigns positions to nodes in such a way that minimizes the forces between them, resulting in a visually appealing layout.
When you call nx.spring_layout(G) without specifying any arguments, the layout is generated randomly each time the function is called. However, if you want to ensure that you get the same layout each time you generate it, you can set a seed for the random number generator.
PYTHON
# Define positions for nodes
# Set a seed for the random number generator
seed_value = 42 # Choose any integer value as the seed
pos = nx.spring_layout(G, seed=seed_value)
posOUTPUT
{1: array([0.4112362 , 0.99648922]),
2: array([ 1. , -0.41570474]),
3: array([-0.41137359, -0.99514183]),
4: array([-0.99986261, 0.41435735])}Try removing or changing the seed_value; what do you observe?
We create traces for edges and nodes. Each edge is represented by a line connecting the positions of its two endpoints, and each node is represented by a marker at its position.
PYTHON
# Create edge traces
edge_traces = []
for edge in G.edges():
x0, y0 = pos[edge[0]]
x1, y1 = pos[edge[1]]
edge_trace = go.Scatter(x=[x0, x1], y=[y0, y1], mode='lines', line=dict(width=3))
edge_traces.append(edge_trace)
edge_tracesOUTPUT
[Scatter({
'line': {'width': 3},
'mode': 'lines',
'x': [0.4112362006825586, 1.0],
'y': [0.9964892191512681, -0.41570474278971886]
}),
Scatter({
'line': {'width': 3},
'mode': 'lines',
'x': [0.4112362006825586, -0.9998626088117056],
'y': [0.9964892191512681, 0.41435735265600393]
}),
Scatter({
'line': {'width': 3},
'mode': 'lines',
'x': [1.0, -0.4113735918708533],
'y': [-0.41570474278971886, -0.9951418290175523]
}),
Scatter({
'line': {'width': 3},
'mode': 'lines',
'x': [-0.4113735918708533, -0.9998626088117056],
'y': [-0.9951418290175523, 0.41435735265600393]
})]
Productos pagados de Colab - Cancela los contratos aquí
We create a Plotly figure with the specified data and layout. We disable the legend for simplicity.
PYTHON
node_x = []
node_y = []
for node in G.nodes():
x, y = pos[node]
node_x.append(x)
node_y.append(y)
node_trace = go.Scatter(x=node_x, y=node_y, mode='markers', marker=dict(size=14, color='rgb(255,0,0)'))
print("node_x",node_x)
print("node_y",node_y)OUTPUT
node_x [0.4112362006825586, 1.0, -0.4113735918708533, -0.9998626088117056]
node_y [0.9964892191512681, -0.41570474278971886, -0.9951418290175523, 0.41435735265600393]Let’s create the Plotly figure and show it using Plotly’s show() method.
PYTHON
fig = go.Figure(data=edge_traces + [node_trace], layout=go.Layout(showlegend=False))
fig.show()In the Topological Data Analysis for Pangenomics lesson, we need to graph and color triangles beside edges and nodes. Let’s do an example with NetworkX. First, we define two triangles, one with vertexes (1,2,3) and another with vertexes in nodes (4,3,2) of the above object.
Now, we will use node_trace to define some characteristics of the nodes in the graph. Each node is represented by a marker (dot) with a corresponding text label displayed above it. The parameters used in its creation control various aspects of the appearance and behavior of the markers and text labels.
x=[] and y=[]: These parameters specify the x and y coordinates of the nodes in the scatter plot. Since we’re defining a trace for nodes, these lists are initially empty. The actual coordinates of the nodes will be added later based on the graph’s layout.
mode=‘markers+text’: This parameter specifies the scatter plot’s mode. markers+text indicates that markers (dots representing nodes) and text labels will be displayed for each node.
hoverinfo=‘text’: This parameter specifies what information will be displayed when hovering over a node. In this case, it’s set to ‘text’, meaning the text labels provided for each node will be displayed when hovering over the corresponding marker.
marker=dict(size=14): This parameter specifies the properties of the markers (nodes) in the scatter plot. Here, size=14 indicates the size of the markers.
text=[‘Node 1’, ‘Node 2’, ‘Node 3’]: This parameter specifies the text labels for each node.
textposition=‘top center’: This parameter specifies the text’s position relative to the markers.
textfont=dict(size=14): This parameter specifies the font properties of the text labels. Here, size=14 indicates the font size of the text.
PYTHON
# Node trace
node_trace = go.Scatter(x=[], y=[], mode='markers+text', hoverinfo='text', marker=dict(size=14), text=['Node 1', 'Node 2', 'Node 3'], textposition='top center', textfont=dict(size=14))Now that you know the for cycle, let’s iterate over all
edges in the graph, extract the positions of the nodes connected by each
edge, and create a scatter plot trace representing the edge with a line
connecting the two endpoints. These traces are stored in a list
(edge_traces) to be later included in the plot.
PYTHON
# Edge traces
edge_traces = []
for edge in G.edges():
x0, y0 = pos[edge[0]]
x1, y1 = pos[edge[1]]
edge_trace = go.Scatter(x=[x0, x1, None], y=[y0, y1, None], mode='lines', line=dict(width=3, color='rgba(0,0,0,0.5)'))
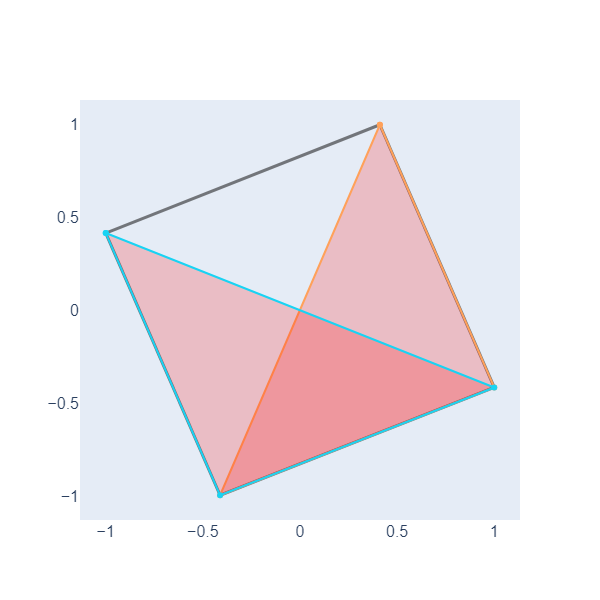
edge_traces.append(edge_trace)Now, we iterate over all triangles in the graph, extract the positions of the vertices of each triangle, and create scatter plot traces representing the triangles as filled polygons with lines connecting the vertices. These traces are stored in a list (triangle_traces) to be later included in the plot.
PYTHON
# Triangle traces
triangle_traces = []
for triangle in triangles:
x = [pos[vertex][0] for vertex in triangle]
y = [pos[vertex][1] for vertex in triangle]
triangle_trace = go.Scatter(x=x + [x[0]], y=y + [y[0]], fill='toself', mode='lines+markers', line=dict(width=2), fillcolor='rgba(255,0,0,0.2)')
triangle_traces.append(triangle_trace)Now, we want to configure the plot layout by specifying settings for various components such as the legend, hover behavior, appearance of axes, and font properties of tick labels. These settings are organized into a layout object (layout) using the go.Layout() constructor.
PYTHON
# Configure the layout of the plot
layout = go.Layout(showlegend=False, hovermode='closest', xaxis=dict(showgrid=False, zeroline=False, tickfont=dict(size=16, family='Arial, sans-serif')), yaxis=dict(showgrid=False, zeroline=False, tickfont=dict(size=16, family='Arial, sans-serif')))To create the figure, we add edges, triangles, and nodes to the data and then use the go.Figure function.
PYTHON
# Create the figure
fig = go.Figure(data=edge_traces + triangle_traces + [node_trace], layout=layout)Finally, we want to adjust the size of the plot by setting its width and height based on the plot_size and dpi variables.
PYTHON
# Set the figure size
plot_size = 1
dpi = 600
fig.update_layout(width=plot_size * dpi, height=plot_size * dpi)
fig.show() # Show the figureHere, we have plotted two filled triangles, we will need this ability to graph objects called simplicial complexes in the following lesson.
Exercise 2(Intermediate): Documenting a graph plot code
Look at the following function used in the Horizontal Gene transfer episode.
Based on what you learned about plotting triangles and edges, sort the comments that document what that part of the code is doing.
COMMENT: Triangle traces
COMMENT: Calculate node positions if not provided
COMMENT: Show the figure
COMMENT: Node trace
COMMENT: Save the figure if a filename is provided
COMMENT: Configure the layout of the plot
COMMENT: Edge tracesWhat do you think the simplex tree contain?
what is the save_fiename doing?
PYTHON
def visualize_simplicial_complex(simplex_tree, filtration_value, vertex_names=None, save_filename=None, plot_size=1, dpi=600, pos=None):
G = nx.Graph()
triangles = [] # List to store triangles (3-nodes simplices)
for simplex, filt in simplex_tree.get_filtration():
if filt <= filtration_value:
if len(simplex) == 2:
G.add_edge(simplex[0], simplex[1])
elif len(simplex) == 1:
G.add_node(simplex[0])
elif len(simplex) == 3:
triangles.append(simplex)
# FIRST COMMENT
if pos is None:
pos = nx.spring_layout(G)
# SECOND COMMENT
x_values, y_values = zip(*[pos[node] for node in G.nodes()])
node_labels = [vertex_names[node] if vertex_names else str(node) for node in G.nodes()]
node_trace = go.Scatter(x=x_values, y=y_values, mode='markers+text', hoverinfo='text', marker=dict(size=14), text=node_labels, textposition='top center', textfont=dict(size=14))
# THIRD COMMENT
edge_traces = []
for edge in G.edges():
x0, y0 = pos[edge[0]]
x1, y1 = pos[edge[1]]
edge_trace = go.Scatter(x=[x0, x1, None], y=[y0, y1, None], mode='lines', line=dict(width=3, color='rgba(0,0,0,0.5)'))
edge_traces.append(edge_trace)
# FOURTH COMMENT
triangle_traces = []
for triangle in triangles:
x0, y0 = pos[triangle[0]]
x1, y1 = pos[triangle[1]]
x2, y2 = pos[triangle[2]]
triangle_trace = go.Scatter(x=[x0, x1, x2, x0, None], y=[y0, y1, y2, y0, None], fill='toself', mode='lines+markers', line=dict(width=2), fillcolor='rgba(255,0,0,0.2)')
triangle_traces.append(triangle_trace)
# 5Th COMMENT
layout = go.Layout(showlegend=False, hovermode='closest', xaxis=dict(showgrid=False, zeroline=False, tickfont=dict(size=16, family='Arial, sans-serif')), yaxis=dict(showgrid=False, zeroline=False, tickfont=dict(size=16, family='Arial, sans-serif')))
fig = go.Figure(data=edge_traces + triangle_traces + [node_trace], layout=layout)
# Set the figure size
fig.update_layout(width=plot_size * dpi, height=plot_size * dpi)
# 6th COMMENT
if save_filename:
pio.write_image(fig, save_filename, width=plot_size * dpi, height=plot_size * dpi, scale=1)
# 7th COMMENT
fig.show()
return GThe sorted comments are:
FIRST COMMENT: Calculate node positions if not provided
SECOND COMMENT: Node trace
THIRD COMMENT: Edge traces
FOURTH COMMENT: Triangle traces
5TH COMMENT: Configure the layout of the plot
6TH COMMENT: Save the figure if a filename is provided
7 COMMENT: Show the figureObject simplex_tree contains information about nodes, edges, and triangles
Is a parameter to save our graphs in a file
- matplotlib is a library
- NetworkX is a library
Content from Plotting
Last updated on 2026-02-20 | Edit this page
Overview
Questions
- How do I create plots in Python?
Objectives
- Create basic plots in Python using the Matplotlib library
There are several different Python plotting libraries. Perhaps the most popular one is Matplotlib, initially released back in 2003. Another library, called Seaborn, is based on Matplotlib, and provides “nicer” defaults for colors and plots, making it easier to build beautiful publication-ready plots. A modern alternative is Plotly, that is available not only in Python, but also in R, Julia, JavaScript, MatLab, and F#. This tutorial serves as a brief introduction into Matplotlib.
Introduction to plots: histograms
We’ll use a dataset of reference genomes of Streptomyces. To download it and load it into the Python environment, we can use the Pandas library.
PYTHON
import pandas as pd
data = pd.read_csv("https://raw.githubusercontent.com/carpentries-incubator/pangenomics-python/gh-pages/data/streptomyces.csv")
dataMatplotlib is a large library; however, most plotting functions are
available in the matplotlib.pyplot module, which is usually
imported as follows.
The Matplotlib official website provides a convenient
page showing the main plot types available. We’ll begin with a
histogram showing the number of genes per reference assembly, which can
be accomplished by using the plt.hist function and pass the
"genes" column of the dataset, grouping the values
automatically. Use the plt.show() function to draw the plot
on the screen.
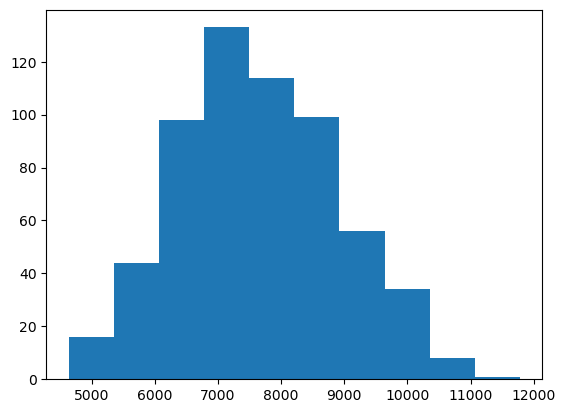
The plt.hist function has many
many options that allow you to customize how the chart looks like.
For instance, we can use the bins parameter to set the
number of bars in the histogram. Furthermore, plots are useless without
proper labels, so we’ll use the plt.xlabel,
plt.ylabel and plt.title functions to define
the label for the x axis, the label for the y axis, and the title for
the plot, respectively.
PYTHON
plt.hist(data["genes"], bins=20)
plt.xlabel("Number of genes")
plt.ylabel("Number of assemblies")
plt.title("Number of genes per Streptomyces assembly")
plt.show()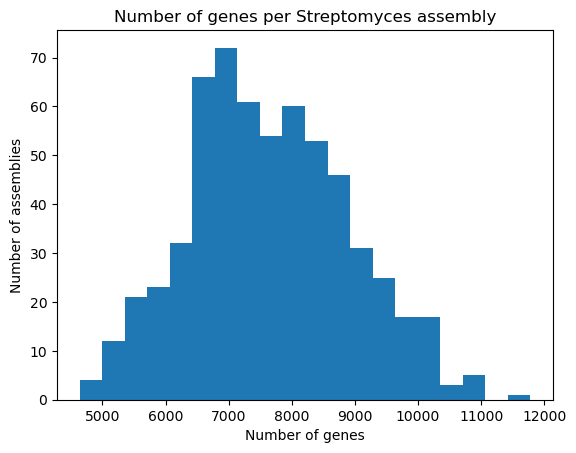
Bars and lines
Whereas plt.hist allows you to pass the variables
directly, other plot types require you to perform some manipulations on
the dataset, because we should explicitly provide both the x and y axes.
The plt.bar function, as its name suggests, creates
bar plots; we’ll use it to visualize the number of chromosomes per
assembly in our dataset. First, we take the "chromosomes"
column from the dataset, and use the .value_counts() method
to count how many times each chromosome count appears in it. This method
returns a Pandas Series, with an index and
values which we can access. So, in order to build the bar
plot, we first provide the unique chromosome counts for the x axis, and
the values for the y axis. We’ll also change the bar colors to dark red
and add labels.
OUTPUT
chromosomes
1.0 100
2.0 42
3.0 15
4.0 3
5.0 3
Name: count, dtype: int64PYTHON
plt.bar(chromosomes.index, chromosomes.values, color="darkred")
plt.xlabel("Number of chromosomes")
plt.ylabel("Number of assemblies")
plt.title("Number of chromosomes per assembly in Streptomyces")
plt.show()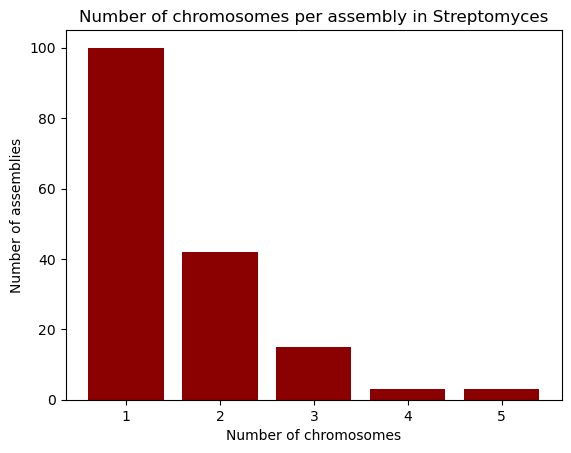
Going horizontal: creating horizontal bar plots
If you wish to use a horizontal bar plot instead of a vertical one,
use the plt.barh function. Don’t forget to change your
labels accordingly!
Let’s now learn how to build line plots using plt.plot
by visualizing the number of reference assemblies released by year.
Similar to the previous example, we’ll use the
.value_counts method on the "release_year"
columns to count the number of assemblies per year; however, this method
sorts the index by the count, so in order to keep the original order
(which is already chronological), we pass the sort=False
parameter. Next, we provide the index and values for the x and y axes of
our plot.
OUTPUT
release_year
2008 1
2009 3
2010 1
2011 1
2012 2
2013 8
2014 35
2015 24
2016 43
2017 25
2018 44
2019 76
2020 153
2021 81
2022 51
2023 34
2024 21
Name: count, dtype: int64PYTHON
plt.plot(genomes_year.index, genomes_year.values)
plt.xlabel("Year")
plt.ylabel("Released assemblies")
plt.title("Released reference Streptomyces assemblies per year")
plt.show()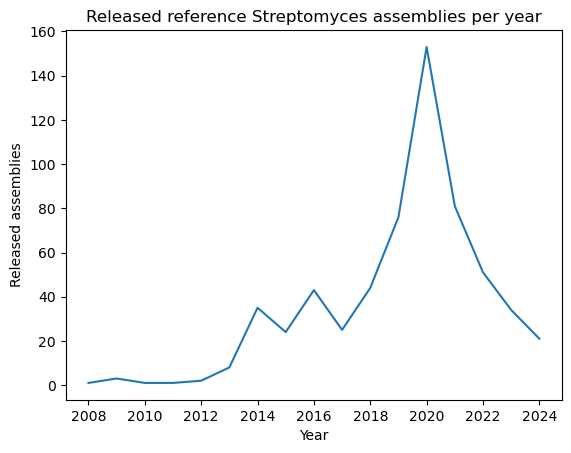
A nice feature about plt.plot is that we can change the
way the line looks like, either by modifying the edges and/or the
vertices. You can find the format guide in the “Format Strings” section
of the function’s
documentation. Some example strings you can use are depicted in the
next code block.
PYTHON
"--" # Dashed line
":" # Dotted line
"o" # Large dots only
"v" # Down-facing triangles only
"s" # Squares only
"--o" # Dashed line with large dots
":s" # Dotted line with squaresLet’s modify our plot by making it dotted with large dots, with a dark green color.
PYTHON
plt.plot(genomes_year.index, genomes_year.values, ':o', color="darkgreen")
plt.xlabel("Year")
plt.ylabel("Released Genomes")
plt.title("Released Reference Streptomyces Genomes per Year")
plt.show()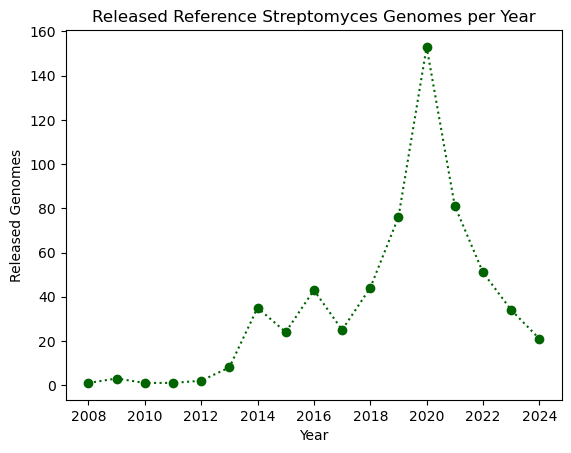
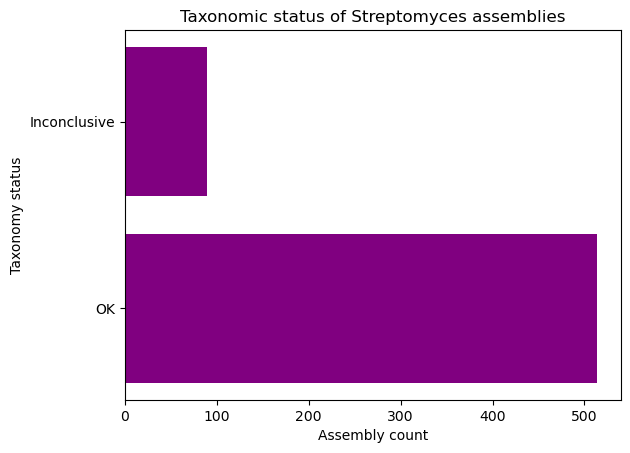
Exercise (Beginner): Plotting with Matplotlib
Complete the following code block to create a horizontal bar plot with the number of assemblies with conclusive and inconclusive taxonomy from the dataset. Use purple to color the bars.
Multiple plots in a single figure
The plt.subplots(x, y)
function creates a multi-plot figure with x rows and
y columns. It returns a Figure object that
allows to modify general aspects of the figure, and an empty array which
will store the plots and are accessible via indices. As an example,
we’ll create a figure with one column and three rows and place the three
plots be made in the lesson. Instead of using .xlabel,
.ylabel and .title, we use
.set_xlabel, .set_ylabel and
.set_title, respectively. At the end, we use the
set_figheight method to set the height for the entire
figure, and the .tight_layout method on the figure in order
to ensure that everything fits in properly.
PYTHON
# Figure initialization
fig, ax = plt.subplots(3)
# First plot: histogram
ax[0].hist(data["genes"], bins=20)
ax[0].set_xlabel("Number of genes")
ax[0].set_ylabel("Number of assembly")
ax[0].set_title("Genes per assembly")
# Second plot: bar
ax[1].bar(chromosomes.index, chromosomes.values, color="darkred")
ax[1].set_xlabel("Number of chromosomes")
ax[1].set_ylabel("Number of assemblies")
ax[1].set_title("Chromosomes per assembly")
# Third plot: line
ax[2].plot(genomes_year.index, genomes_year.values, ':o', color="darkgreen")
ax[2].set_xlabel("Year")
ax[2].set_ylabel("Released Genomes")
ax[2].set_title("Assemblies per year")
# Figure configuration
fig.set_figheight(12)
fig.tight_layout()
plt.show()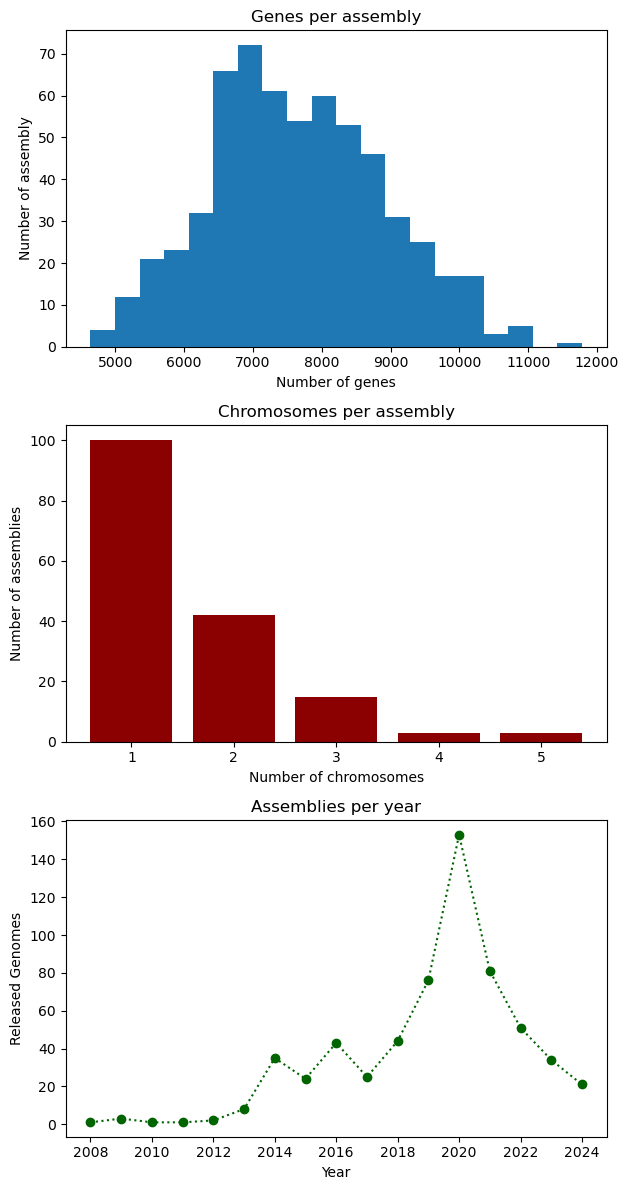
- Matplotlib is a popular plotting library for Python